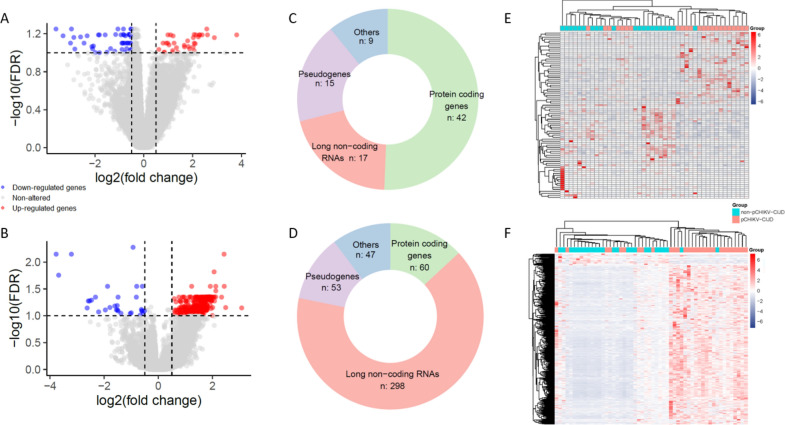
Transcriptomic insights into early mechanisms underlying post-chikungunya chronic inflammatory joint disease.
Chikungunya virus (CHIKV) infection often results in a chronic joint condition known as Post-Chikungunya Chronic Inflammatory Joint Disease (pCHIKV-CIJD). This condition disrupts individuals' daily lives and contributes to increased healthcare expenditure. This study investigated the molecular mechanisms underlying pCHIKV-CIJD development by analyzing RNA transcripts, including small RNAs, of whole blood from CHIKV-infected patients. By comparing patients who evolved to pCHIKV-CIJD with those who did not, we identified molecular signatures associated with chronification in acute and post-acute disease phases. These molecules were primarily associated with an altered immune response regulation. Notably, LIFR, an immune receptor that enhanced IL-6 transcription, was down-regulated in the acute phase of pCHIKV-CIJD patients, while its inhibitor, hsa-miR-98-5p, was up-regulated in these individuals. Other downregulated genes include members of immune mechanisms whose impairment can lead to a reduction in the first line of antiviral response, thereby promoting virus persistence for a longer period in these patients. Additionally, pCHIKV-CIJD patients exhibited reduced transcript levels of MMP8, LFT, and DDIT4, genes already implicated in the pathological process of other types of inflammatory arthritis and seemingly relevant for pCHIKV-CIJD development. Overall, our findings provide insights into the early molecular mechanisms involved in the chronification and highlight potential targets for further investigation.
Authors
Ramundo MS, da Fonseca GC, Ten-Caten F, Gerber AL, et al.
External link
Publication Year
Publication Journal
Associeted Project
Systems Immunology of Human Diseases
Lista de serviços
-
StructRNAfinder: an automated pipeline and web server for RNA families prediction.StructRNAfinder: an automated pipeline and web server for RNA families prediction.
-
CEMiTool: a Bioconductor package for performing comprehensive modular co-expression analyses.CEMiTool: a Bioconductor package for performing comprehensive modular co-expression analyses.
-
webCEMiTool: Co-expression Modular Analysis Made Easy.webCEMiTool: Co-expression Modular Analysis Made Easy.
-
Assessing the Impact of Sample Heterogeneity on Transcriptome Analysis of Human Diseases Using MDP Webtool.Assessing the Impact of Sample Heterogeneity on Transcriptome Analysis of Human Diseases Using MDP Webtool.
-
Predicting RNA Families in Nucleotide Sequences Using StructRNAfinder.Predicting RNA Families in Nucleotide Sequences Using StructRNAfinder.
-
OUTBREAK: a user-friendly georeferencing online tool for disease surveillance.OUTBREAK: a user-friendly georeferencing online tool for disease surveillance.
-
Noninvasive prenatal paternity determination using microhaplotypes: a pilot study.Noninvasive prenatal paternity determination using microhaplotypes: a pilot study.
-
Editorial: User-Friendly Tools Applied to Genetics or Systems Biology.Editorial: User-Friendly Tools Applied to Genetics or Systems Biology.
-
Automatic detection of the parasite Trypanosoma cruzi in blood smears using a machine learning approach applied to mobile phone imagesAutomatic detection of the parasite Trypanosoma cruzi in blood smears using a machine learning approach applied to mobile phone images
-
Tucuxi-BLAST: Enabling fast and accurate record linkage of large-scale health-related administrative databases through a DNA-encoded approachTucuxi-BLAST: Enabling fast and accurate record linkage of large-scale health-related administrative databases through a DNA-encoded approach
-
Ten quick tips for harnessing the power of ChatGPT in computational biologyTen quick tips for harnessing the power of ChatGPT in computational biology