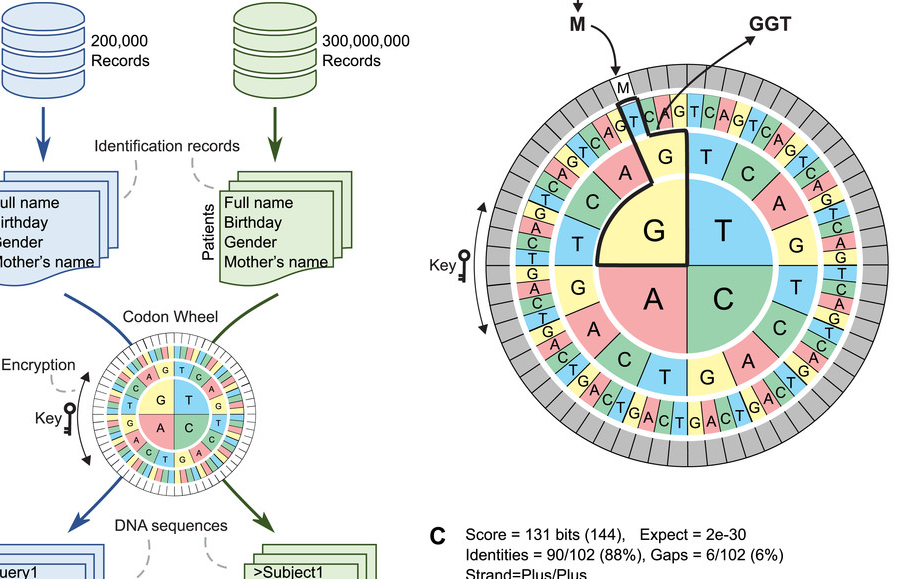
Tucuxi-BLAST: Enabling fast and accurate record linkage of large-scale health-related administrative databases through a DNA-encoded approach
Background. Public health research frequently requires the integration of information from different data sources. However, errors in the records and the high computational costs involved make linking large administrative databases using record linkage (RL) methodologies a major challenge. Methods. We present Tucuxi-BLAST, a versatile tool for probabilistic RL that utilizes a DNA-encoded approach to encrypt, analyze and link massive administrative databases. Tucuxi-BLAST encodes the identification records into DNA. BLASTn algorithm is then used to align the sequences between databases. We tested and benchmarked on a simulated database containing records for 300 million individuals and also on four large administrative databases containing real data on Brazilian patients. Results. Our method was able to overcome misspellings and typographical errors in administrative databases. In processing the RL of the largest simulated dataset (200k records), the state-of-the-art method took 5 days and 7 h to perform the RL, while Tucuxi-BLAST only took 23 h. When compared with five existing RL tools applied to a gold-standard dataset from real health-related databases, Tucuxi-BLAST had the highest accuracy and speed. By repurposing genomic tools, Tucuxi-BLAST can improve data-driven medical research and provide a fast and accurate way to link individual information across several administrative databases.
Authors
Araujo, Jose Deney; Santos-e-Silva, Juan Carlo; Costa-Martins, Andre Guilherme; Sampaio, Vanderson; de Castro, Daniel Barros; de Souza, Robson F; Giddaluru, Jeevan; Ramos, Pablo Ivan P; Pita, Robespierre; Barreto, Mauricio L;
External link
Publication Year
Publication Journal
Associeted Project
User-friendly computational Tools
Lista de serviços
-
As antisense RNA gets intronic.As antisense RNA gets intronic.
-
Androgen responsive intronic non-coding RNAs.Androgen responsive intronic non-coding RNAs.
-
Conserved tissue expression signatures of intronic noncoding RNAs transcribed from human and mouse loci.Conserved tissue expression signatures of intronic noncoding RNAs transcribed from human and mouse loci.
-
The intronic long noncoding RNA ANRASSF1 recruits PRC2 to the RASSF1A promoter, reducing the expression of RASSF1A and increasing cell proliferation.The intronic long noncoding RNA ANRASSF1 recruits PRC2 to the RASSF1A promoter, reducing the expression of RASSF1A and increasing cell proliferation.
-
Antisense intronic non-coding RNA levels correlate to the degree of tumor differentiation in prostate cancer.Antisense intronic non-coding RNA levels correlate to the degree of tumor differentiation in prostate cancer.
-
Insight Into the Long Noncoding RNA and mRNA Coexpression Profile in the Human Blood Transcriptome Upon Leishmania infantum Infection.Insight Into the Long Noncoding RNA and mRNA Coexpression Profile in the Human Blood Transcriptome Upon Leishmania infantum Infection.
-
Long non-coding RNAs associated with infection and vaccine-induced immunityLong non-coding RNAs associated with infection and vaccine-induced immunity
-
Comparative transcriptomic analysis of long noncoding RNAs in Leishmania-infected human macrophagesComparative transcriptomic analysis of long noncoding RNAs in Leishmania-infected human macrophages
-
SARS-CoV-2 Selectively Induces the Expression of Unproductive Splicing Isoforms of Interferon, Class I MHC, and Splicing Machinery Genes.SARS-CoV-2 Selectively Induces the Expression of Unproductive Splicing Isoforms of Interferon, Class I MHC, and Splicing Machinery Genes.