Noninvasive prenatal paternity determination using microhaplotypes: a pilot study.
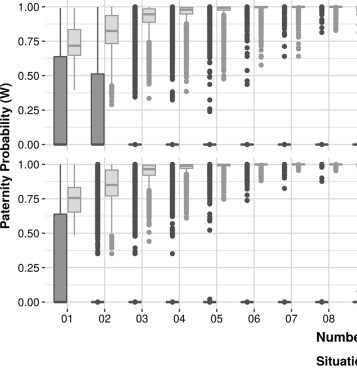
The use of noninvasive techniques to determine paternity prenatally is increasing because it reduces the risks associated with invasive procedures. Current methods, based on SNPs, use the analysis of at least 148 markers, on average. To reduce the number of regions, we used microhaplotypes, which are chromosomal segments smaller than 200 bp containing two or more SNPs. Our method employs massively parallel sequencing and analysis of microhaplotypes as genetic markers. We tested 20 microhaplotypes and ascertained that 19 obey Hardy-Weinberg equilibrium and are independent, and data from the 1000 Genomes Project were used for population frequency and simulations. We performed simulations of true and false paternity, using the 1000 Genomes Project data, to confirm if the microhaplotypes could be used as genetic markers. We observed that at least 13 microhaplotypes should be used to decrease the chances of false positives. Then, we applied the method in 31 trios, and it was able to correctly assign the fatherhood in cases where the alleged father was the real father, excluding the inconclusive results. We also cross evaluated the mother-plasma duos with the alleged fathers for false inclusions within our data, and we observed that the use of at least 15 microhaplotypes in real data also decreases the false inclusions. In this work, we demonstrated that microhaplotypes can be used to determine prenatal paternity by using only 15 regions and with admixtures of DNA.
Authors
Jaqueline Yu Ting Wang; Martin R Whittle; Renato David Puga; Anatoly Yambartsev; André Fujita; Helder I Nakaya
External link
Publication Year
Publication Journal
Associeted Project
User-friendly computational Tools
Lista de serviços
-
As antisense RNA gets intronic.As antisense RNA gets intronic.
-
Androgen responsive intronic non-coding RNAs.Androgen responsive intronic non-coding RNAs.
-
Conserved tissue expression signatures of intronic noncoding RNAs transcribed from human and mouse loci.Conserved tissue expression signatures of intronic noncoding RNAs transcribed from human and mouse loci.
-
The intronic long noncoding RNA ANRASSF1 recruits PRC2 to the RASSF1A promoter, reducing the expression of RASSF1A and increasing cell proliferation.The intronic long noncoding RNA ANRASSF1 recruits PRC2 to the RASSF1A promoter, reducing the expression of RASSF1A and increasing cell proliferation.
-
Antisense intronic non-coding RNA levels correlate to the degree of tumor differentiation in prostate cancer.Antisense intronic non-coding RNA levels correlate to the degree of tumor differentiation in prostate cancer.
-
Insight Into the Long Noncoding RNA and mRNA Coexpression Profile in the Human Blood Transcriptome Upon Leishmania infantum Infection.Insight Into the Long Noncoding RNA and mRNA Coexpression Profile in the Human Blood Transcriptome Upon Leishmania infantum Infection.
-
Long non-coding RNAs associated with infection and vaccine-induced immunityLong non-coding RNAs associated with infection and vaccine-induced immunity
-
Comparative transcriptomic analysis of long noncoding RNAs in Leishmania-infected human macrophagesComparative transcriptomic analysis of long noncoding RNAs in Leishmania-infected human macrophages
-
SARS-CoV-2 Selectively Induces the Expression of Unproductive Splicing Isoforms of Interferon, Class I MHC, and Splicing Machinery Genes.SARS-CoV-2 Selectively Induces the Expression of Unproductive Splicing Isoforms of Interferon, Class I MHC, and Splicing Machinery Genes.